|
7/25/2023 0 Comments Amplifx in silico pcr triangle Since the outbreak started, the World Health Organization (WHO) released some SARS-CoV-2 polymerase chain reaction (PCR) protocol assays produced by different reference institutions in the world 1. Identifying viral genetic material using the PCR technique is considered the gold standard for determining SARS-CoV-2 in nasal swab samples from symptomatic patients. The availability of these primer and probe sequences will make it possible to validate more efficient protocols for identifying SARS-CoV-2. In silico predictions also demonstrated that those primers do not bind to nonspecific targets for human, bacterial, fungal, apicomplexan, and other Betacoronaviruses and less pathogenic sub-strains of coronavirus. We obtained nine systems (forward primer + reverse primer + probe) that potentially anneal to highly conserved regions of the virus genome from these analyses. We subsequently validated those primers and probes in 211,833 SARS-CoV-2 whole-genome sequences. 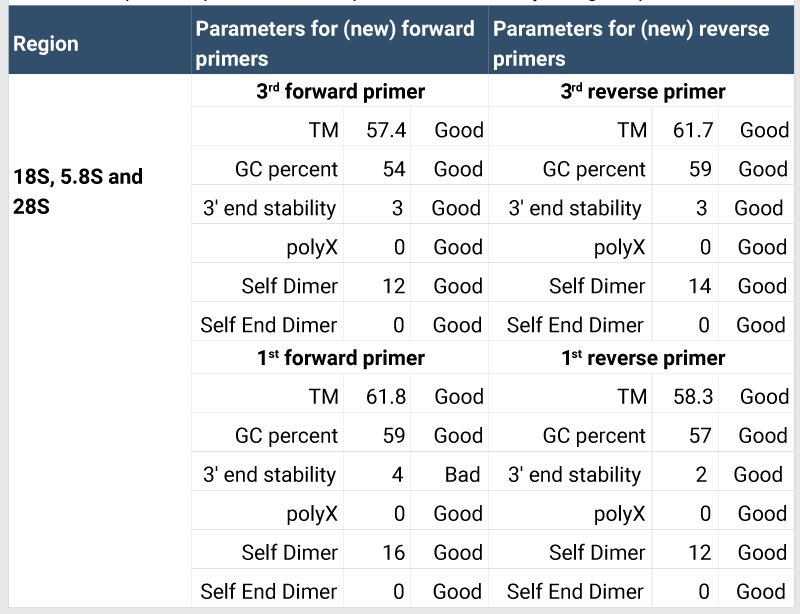
In this work, we designed and described a set of primers and probes targeting conserved regions identified from a multiple sequence alignment of 2341 Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) genomes from the Global Initiative on Sharing All Influenza Data (GISAID). 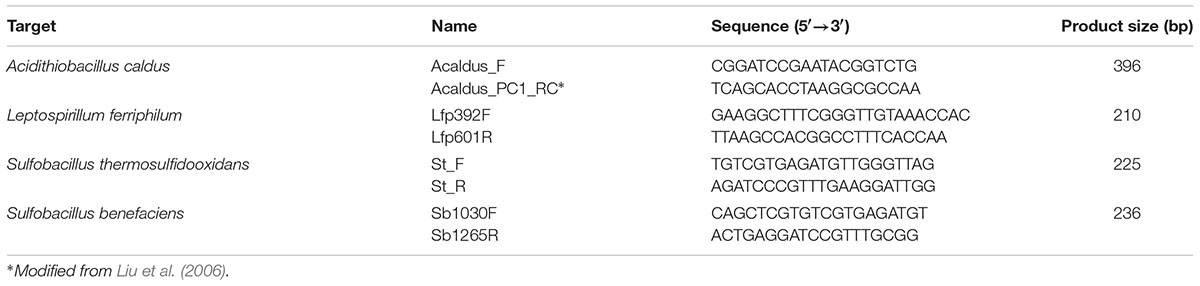
Accurate designing of polymerase chain reaction (PCR) primers targeting conserved segments in viral genomes is desirable for preventing false-negative results and decreasing the need for standardization across different PCR protocols. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
